Fishes | Free Full-Text | Biomass Quantification of the Critically Endangered European eel from Running Waters Using Environmental DNA
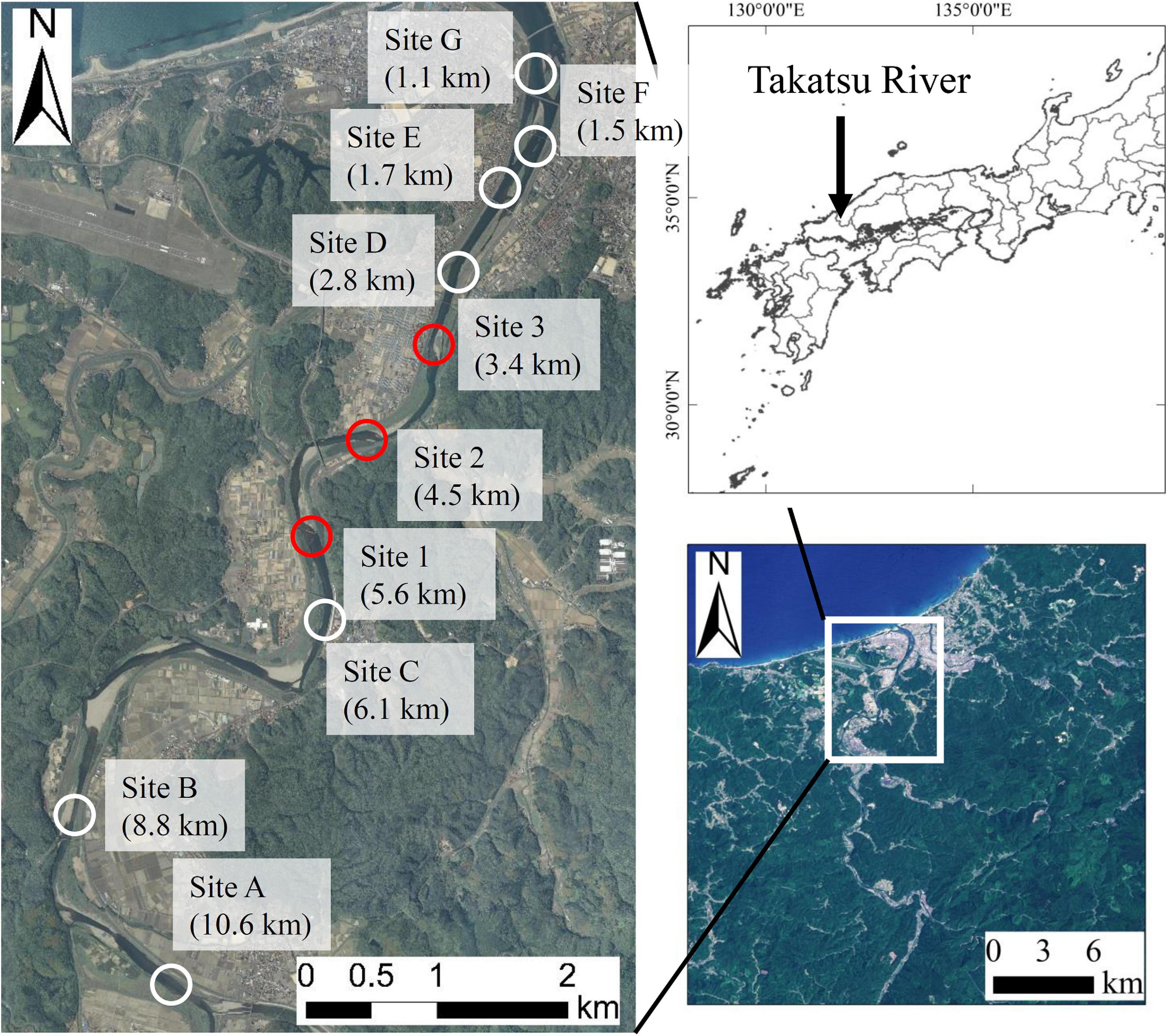
Frontiers | Spatiotemporal Changes of the Environmental DNA Concentrations of Amphidromous Fish Plecoglossus altivelis altivelis in the Spawning Grounds in the Takatsu River, Western Japan

COPMAN: A novel high-throughput and highly sensitive method to detect viral nucleic acids including SARS-CoV-2 RNA in wastewater - ScienceDirect

Environmental DNA reveals the genetic diversity and population structure of an invasive species in the Laurentian Great Lakes | PNAS

Detection of M. tuberculosis in the environment as a tool for identifying high-risk locations for tuberculosis transmission | medRxiv

First detection of critically endangered scalloped hammerhead sharks (Sphyrna lewini) in Guam, Micronesia, in five decades using environmental DNA - ScienceDirect

Detection of M. tuberculosis in the environment as a tool for identifying high-risk locations for tuberculosis transmission | medRxiv

TaqMan-Based Duplex Real-Time PCR Approach for Analysis of Grain Composition (Zea mays - Sorghum bicolor) in Feedstock Flour Mixes for Bioethanol Production | ACS Agricultural Science & Technology

Analysis of the persistence and particle size distributional shift of sperm-derived environmental DNA to monitor Jack Mackerel spawning activity | bioRxiv

Comparison of inhibition resistance among PCR reagents for detection and quantification of environmental DNA - Uchii - 2019 - Environmental DNA - Wiley Online Library

Comparison of inhibition resistance among PCR reagents for detection and quantification of environmental DNA - Uchii - 2019 - Environmental DNA - Wiley Online Library
Spatiotemporal distribution of juvenile chum salmon in Otsuchi Bay, Iwate, Japan, inferred from environmental DNA | PLOS ONE

Field application of an improved protocol for environmental DNA extraction, purification, and measurement using Sterivex filter | Scientific Reports
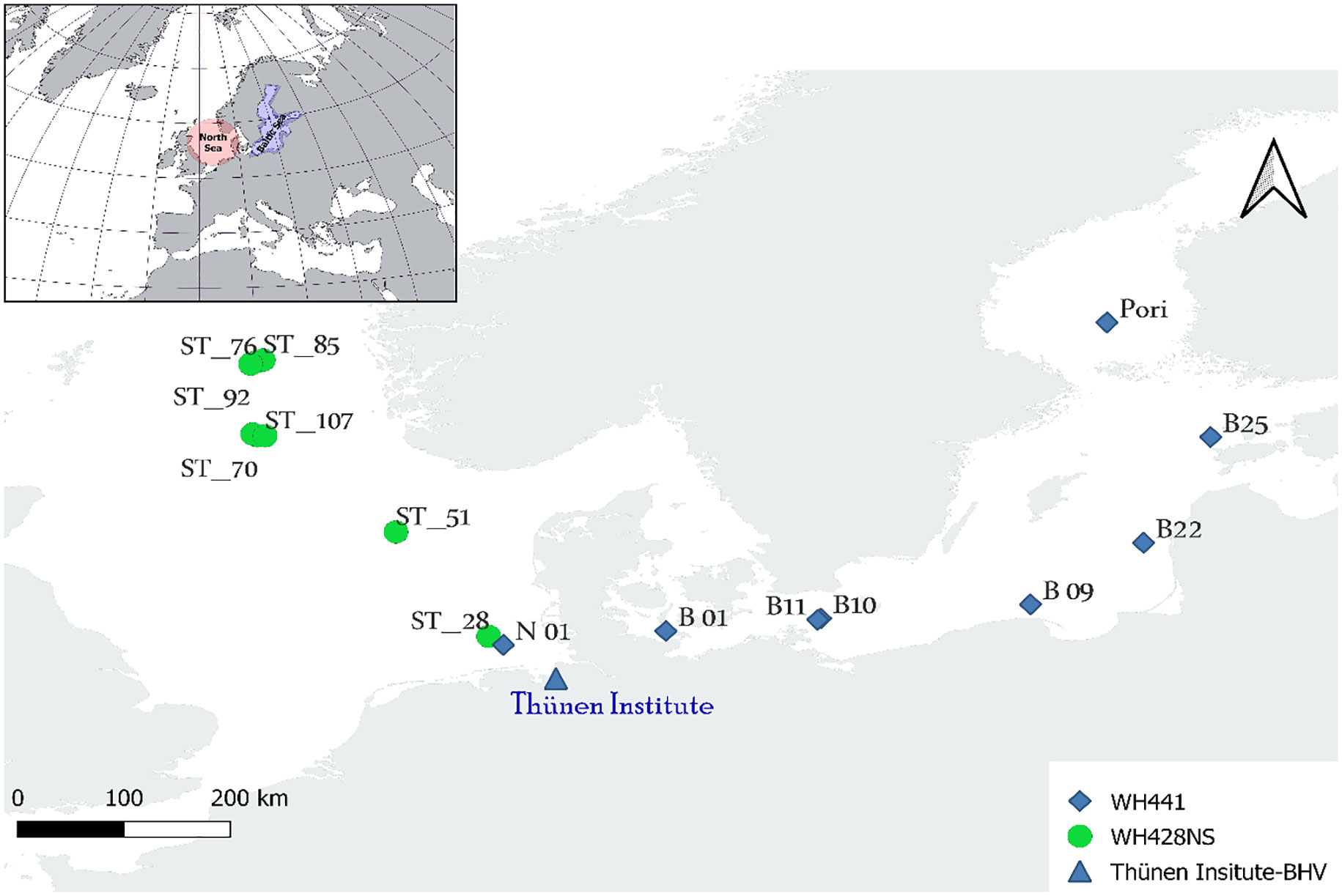
Frontiers | Atlantic cod (Gadus morhua) assessment approaches in the North and Baltic Sea: A comparison of environmental DNA analysis versus bottom trawl sampling

Applied Biosystems™ TaqMan™ Environmental Master Mix 2.0 400 Reactions Applied Biosystems™ TaqMan™ Environmental Master Mix 2.0 | Fisher Scientific

Field application of an improved protocol for environmental DNA extraction, purification, and measurement using Sterivex filter | Scientific Reports

Comparison of inhibition resistance among PCR reagents for detection and quantification of environmental DNA - Uchii - 2019 - Environmental DNA - Wiley Online Library




/placeholder.jpg)